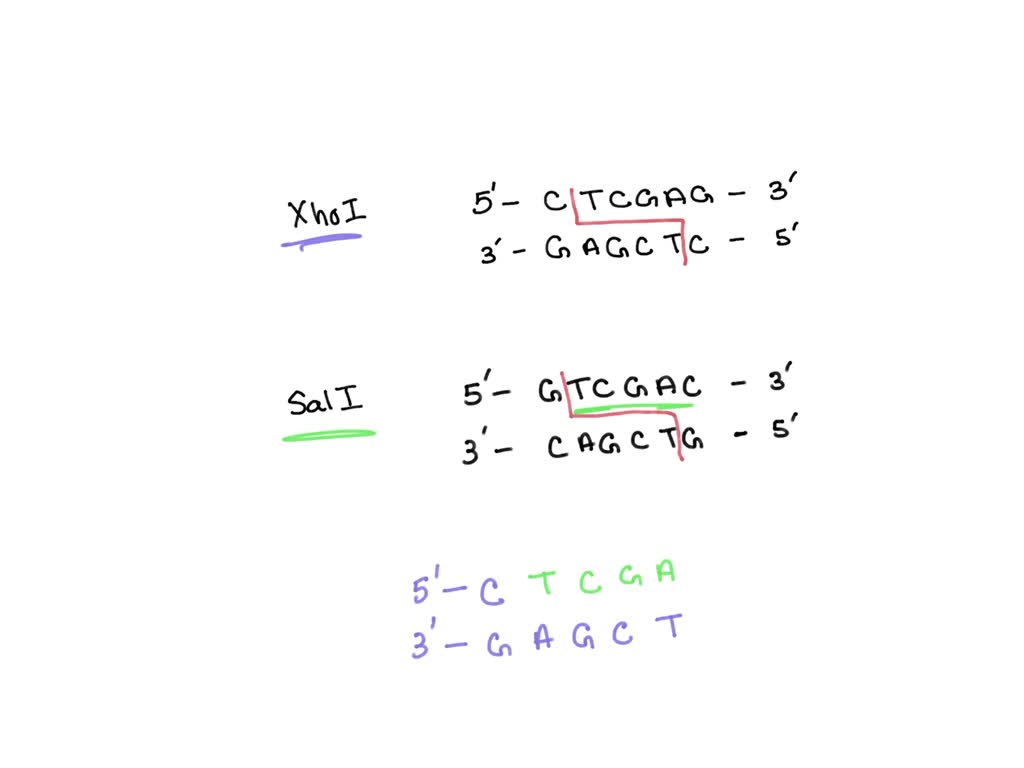
SOLVED: #16) The restriction enzymes Xhol and SalI cut their specific sequences as shown below: XhoI 5' C | TCGAG 3' SalI | 5' GTCGAC 3' 3' GAGC | TC 5' 3'

Digestion of pcEG95-IR plasmid by EcoRI and XhoI restriction enzymes.... | Download Scientific Diagram

A) Restriction enzyme digestion with Xho I and Nco I of the constructs... | Download Scientific Diagram
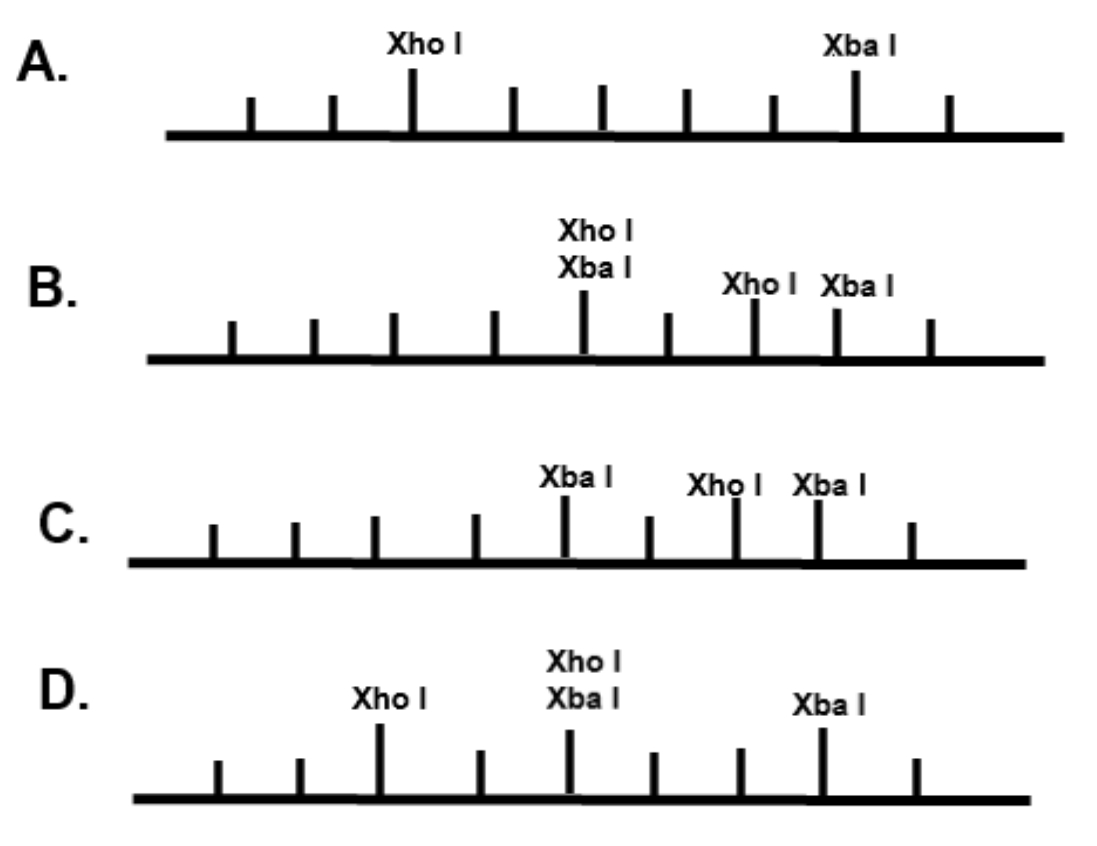
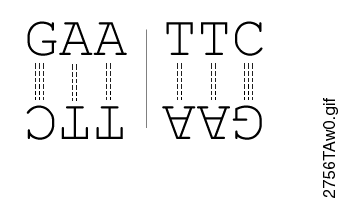
The restriction enzyme XhoI and XbaI were used to digest a linear fragment of DNA at a specific sequence such as that shown for EcoR1 (figure above). Use the information gained from

SciELO - Brasil - In-Situ Gel-Free Plasmid Reassembling for Rapid Gene Subcloning and Truncation In-Situ Gel-Free Plasmid Reassembling for Rapid Gene Subcloning and Truncation