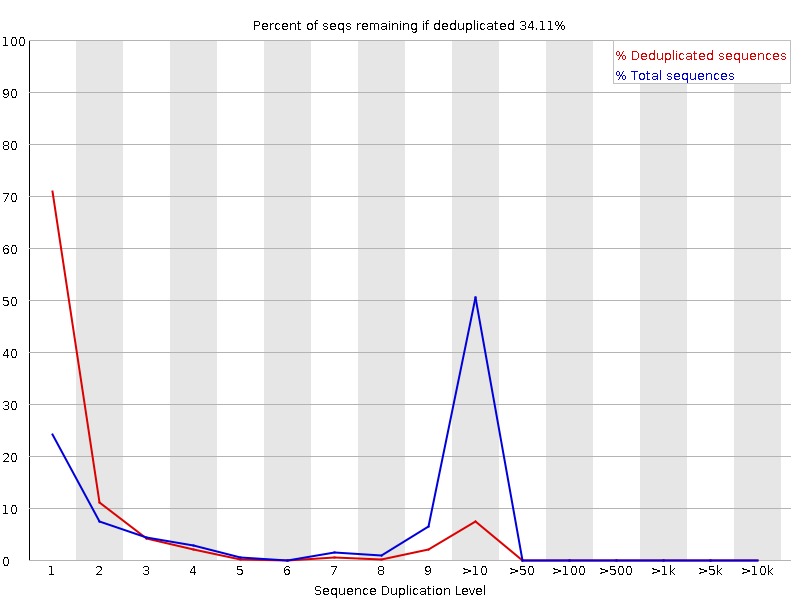
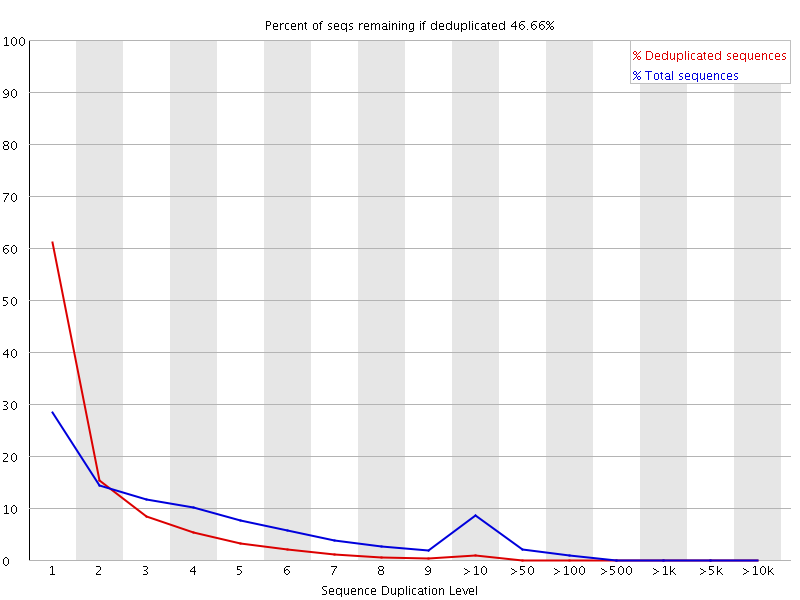
FastQC for RNA-Seq-Fails for per base seq content, gc content, duplication levels, adapter content and kmer content? | ResearchGate
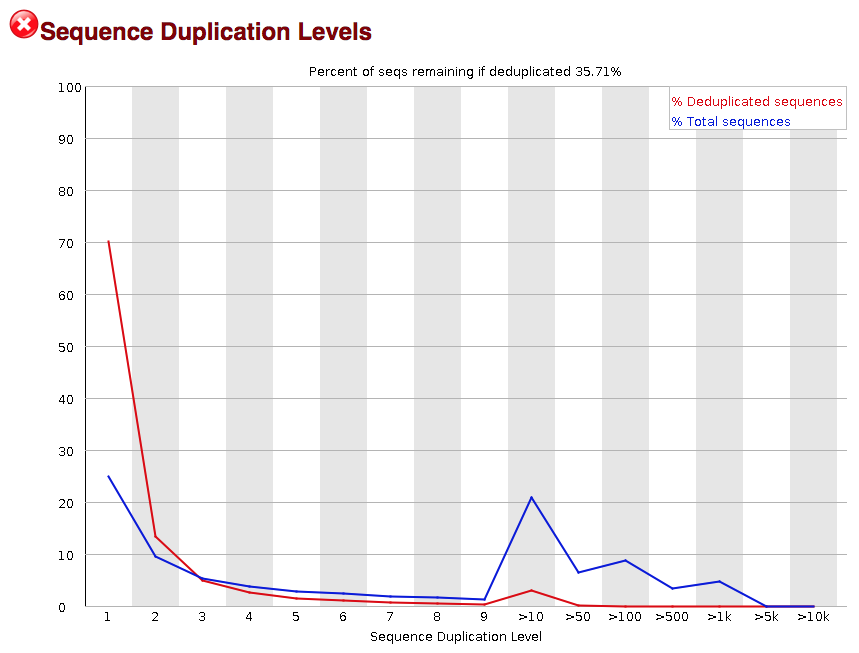
Why does FASTQC show unexpectedly high sequence duplication levels (PCR-duplicates)? | DNA Technologies Core

Summary of FASTQC output for first sample in dataset, Wistar 1 mate 1... | Download Scientific Diagram
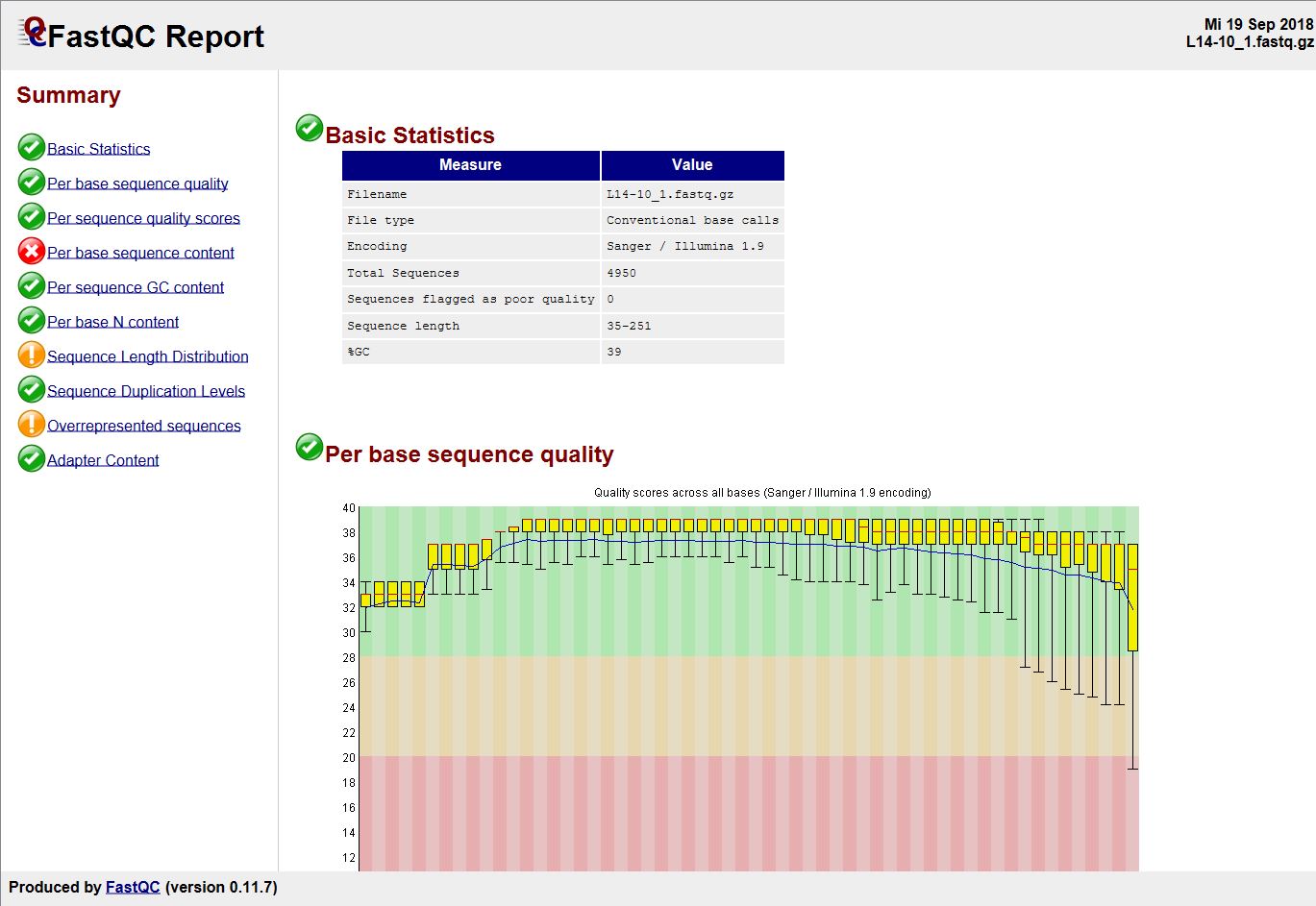
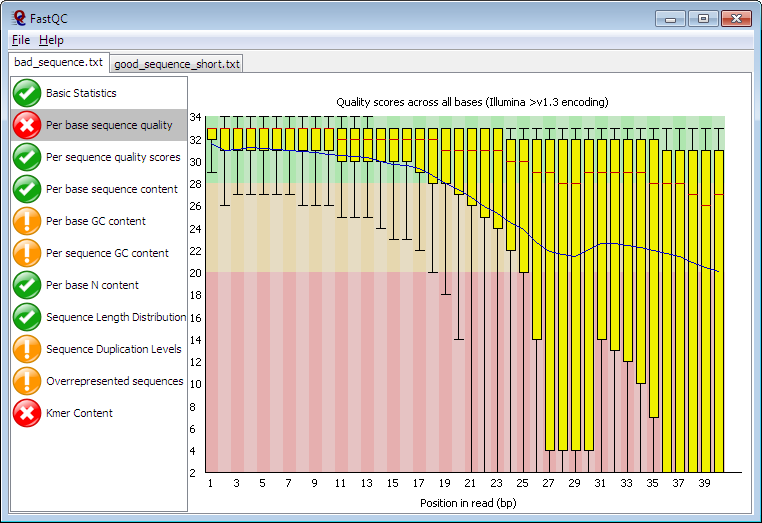
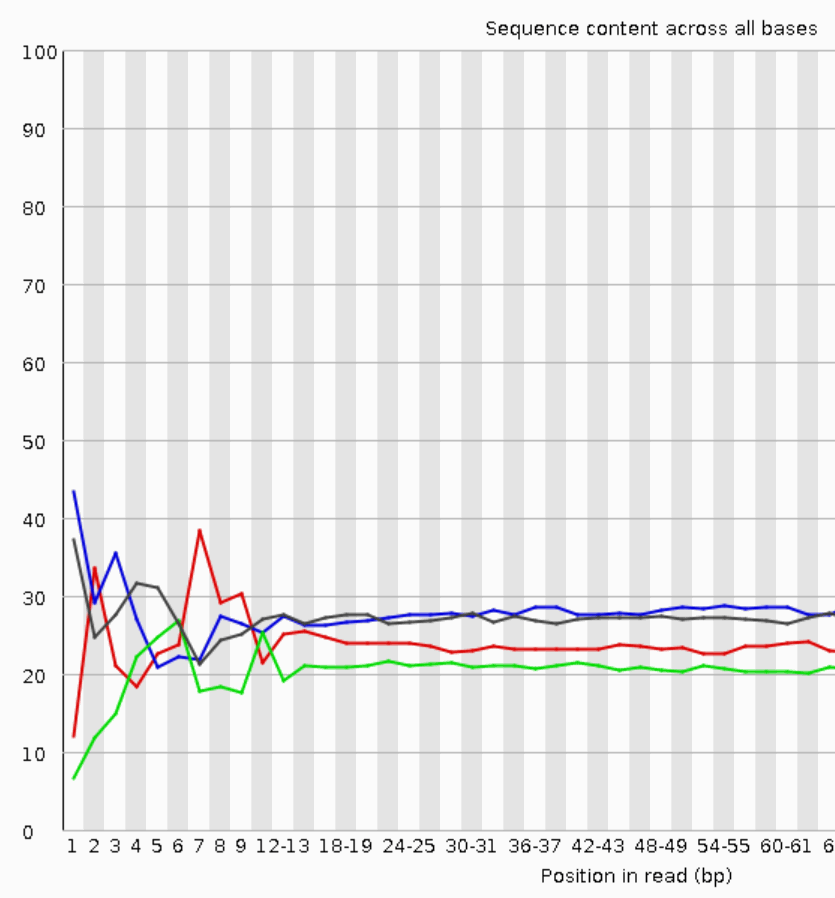
Quality control: Assessing FASTQC results | Introduction to RNA-Seq using high-performance computing - ARCHIVED

FASTQ Quality Assurance Tools - Bioinformatics Team (BioITeam) at the University of Texas - UT Austin Wikis

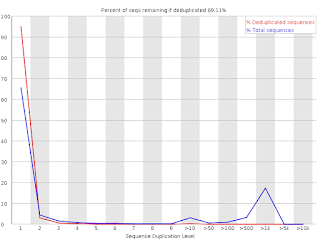
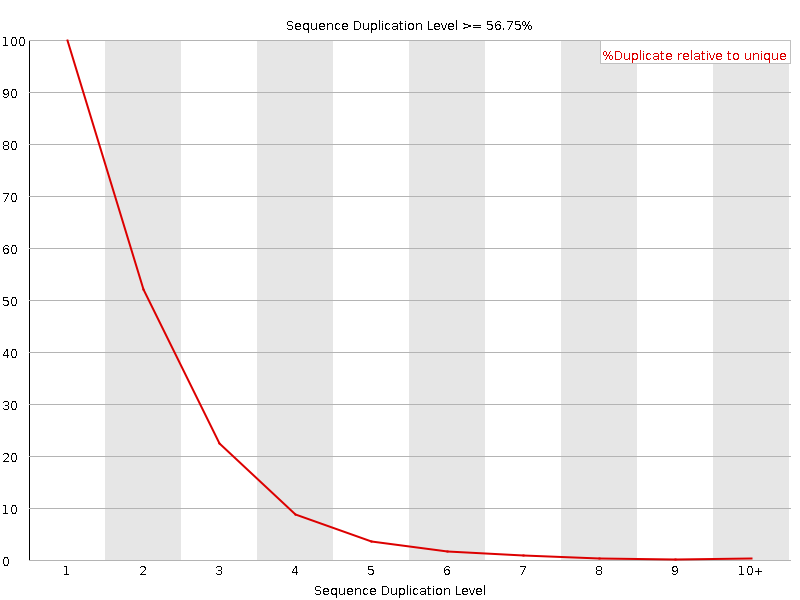
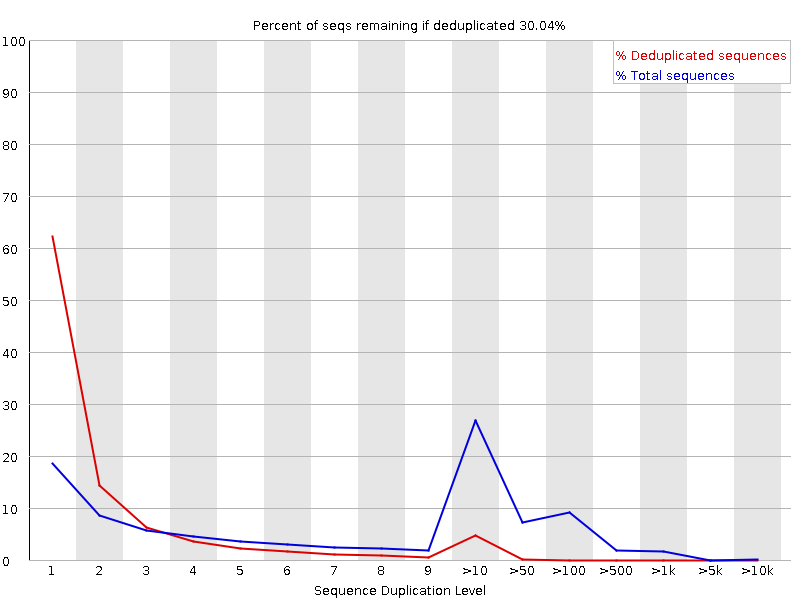
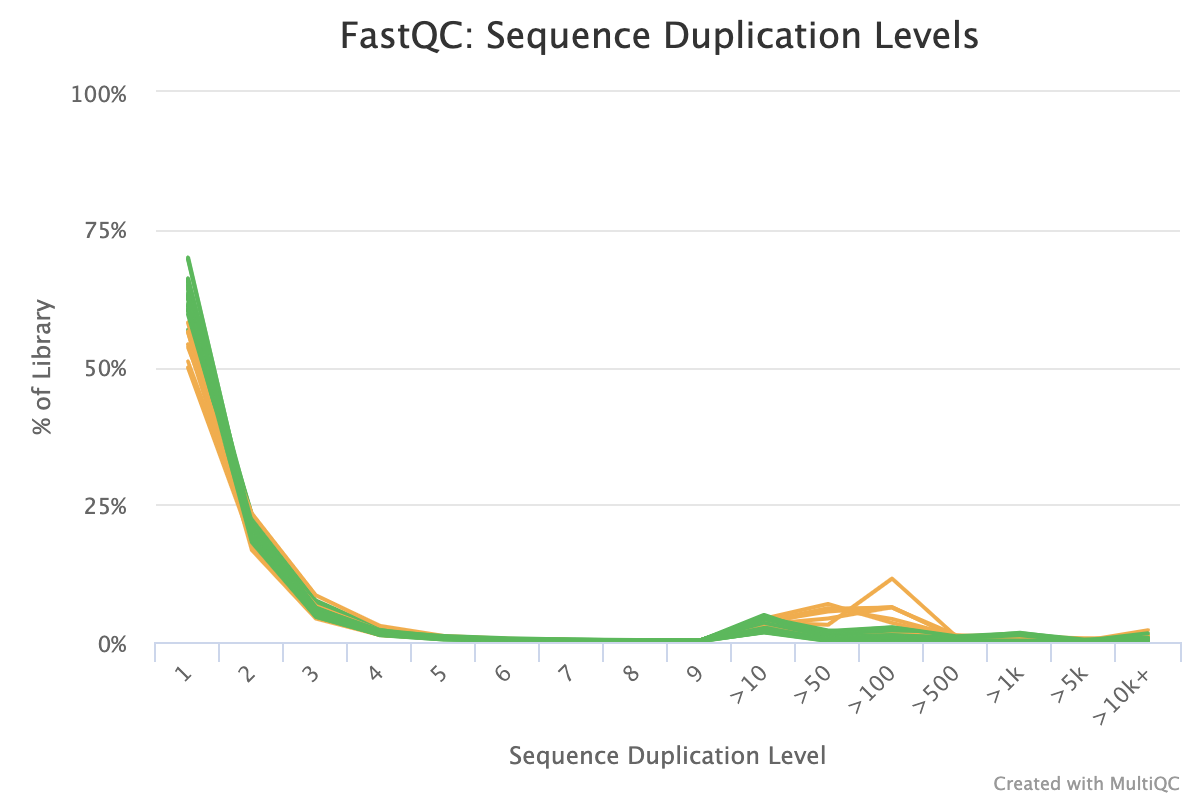
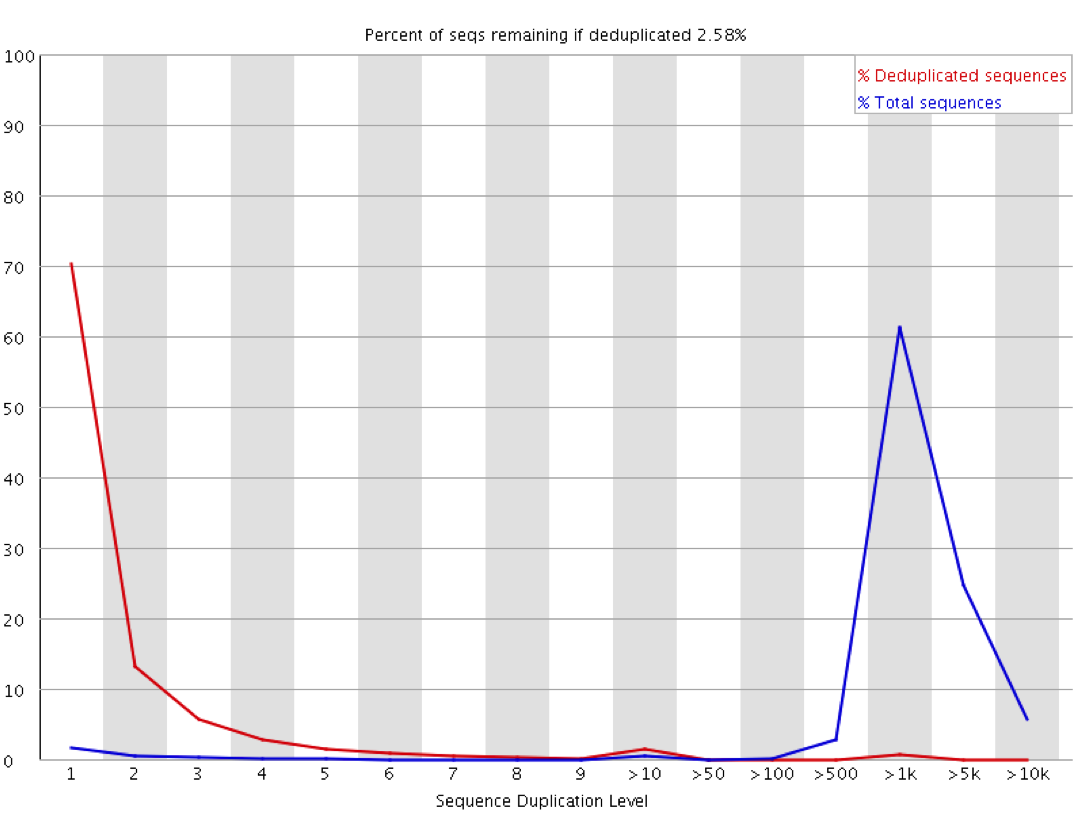
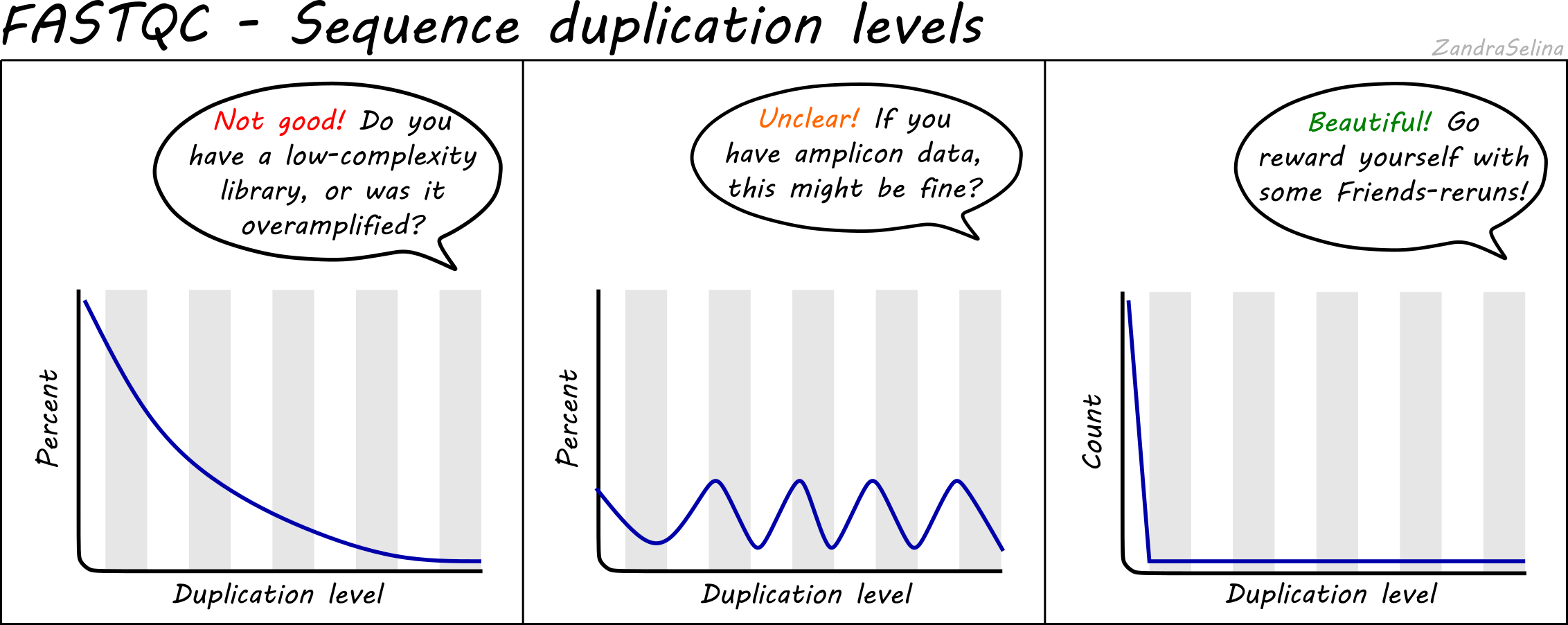
SciBear🇺🇦 on Twitter: "(6/7) Sequence duplication levels. This module counts the degree of #duplication for every sequence in the set and creates a plot showing the relative number of sequences with different

FastQC: Sequence Duplication Levels interactive dialog indexing error · Issue #1092 · ewels/MultiQC · GitHub