BLAST: Compare & identify sequences - NCBI Bioinformatics Resources: An Introduction - Library Guides at UC Berkeley
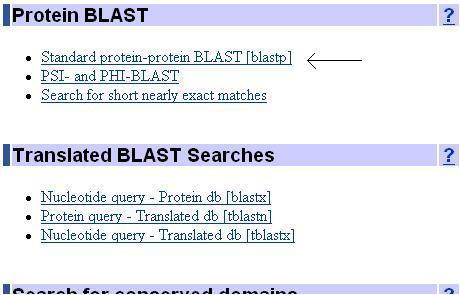
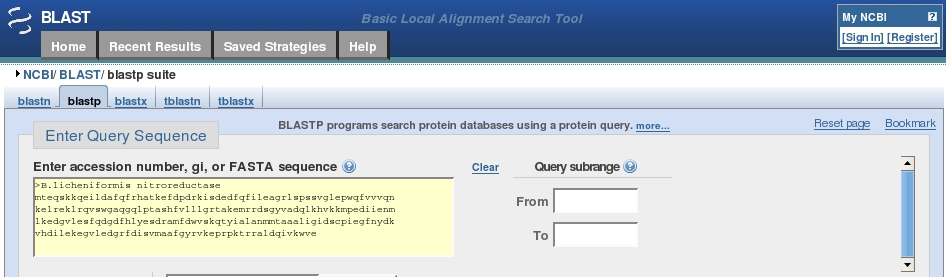
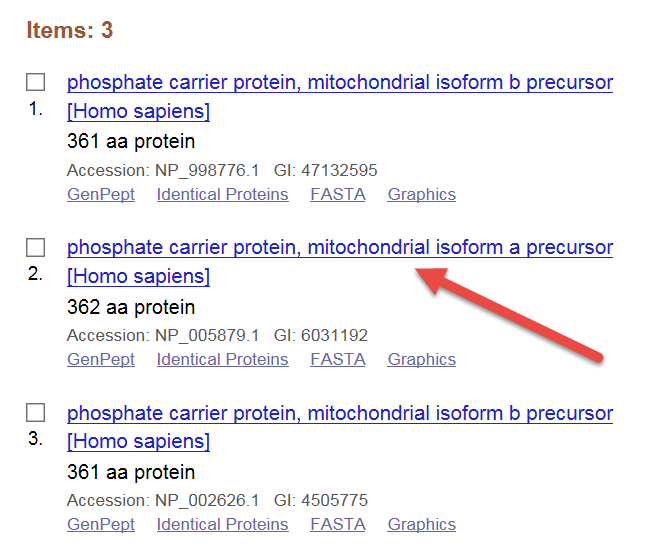
Protein BLAST allows one to input protein sequences and compare these against other protein sequences

BLASTP Protein Sequence Search - Introduction to NCBI Bioinformatics Resources - Learning Resource Center at Uniformed Services University of the Health Sciences

BLASTP Protein Sequence Search - Introduction to NCBI Bioinformatics Resources - Learning Resource Center at Uniformed Services University of the Health Sciences
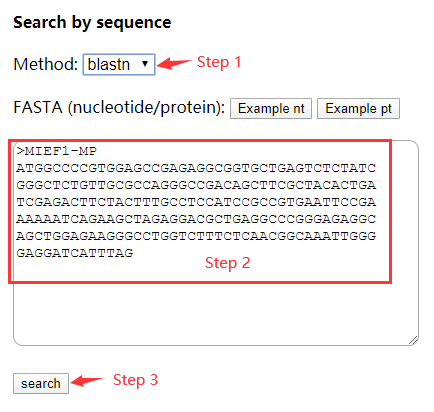
Chapter 5 Workshop 4: Discovering a new gene in a DNA sequence (Wednesday morning) | SWBio Bioinformatics course task book

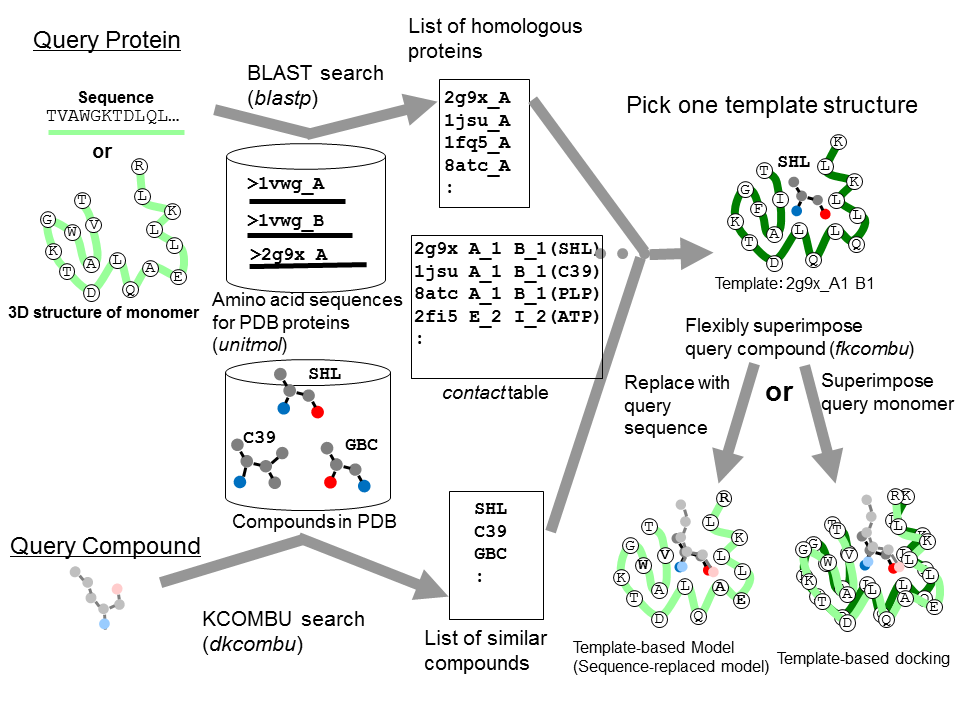
MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets | Nature Biotechnology

The protein BLAST (BLASTp) search against the annotated NCBI protein... | Download Scientific Diagram