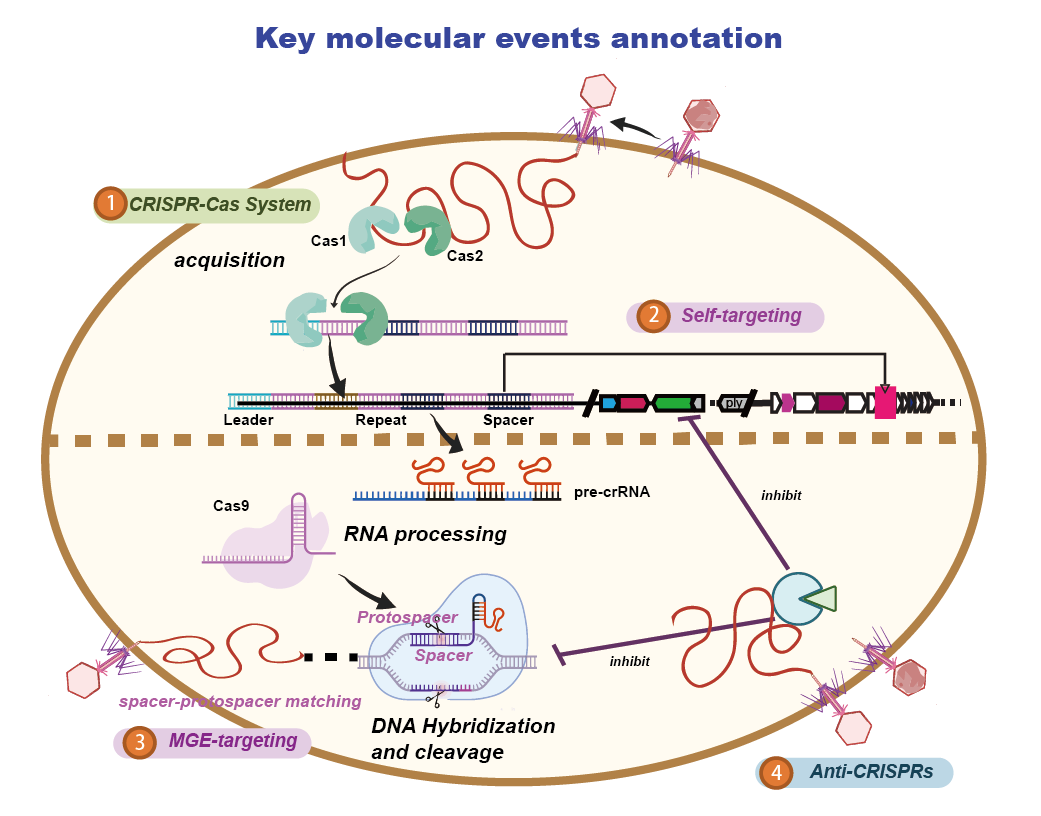
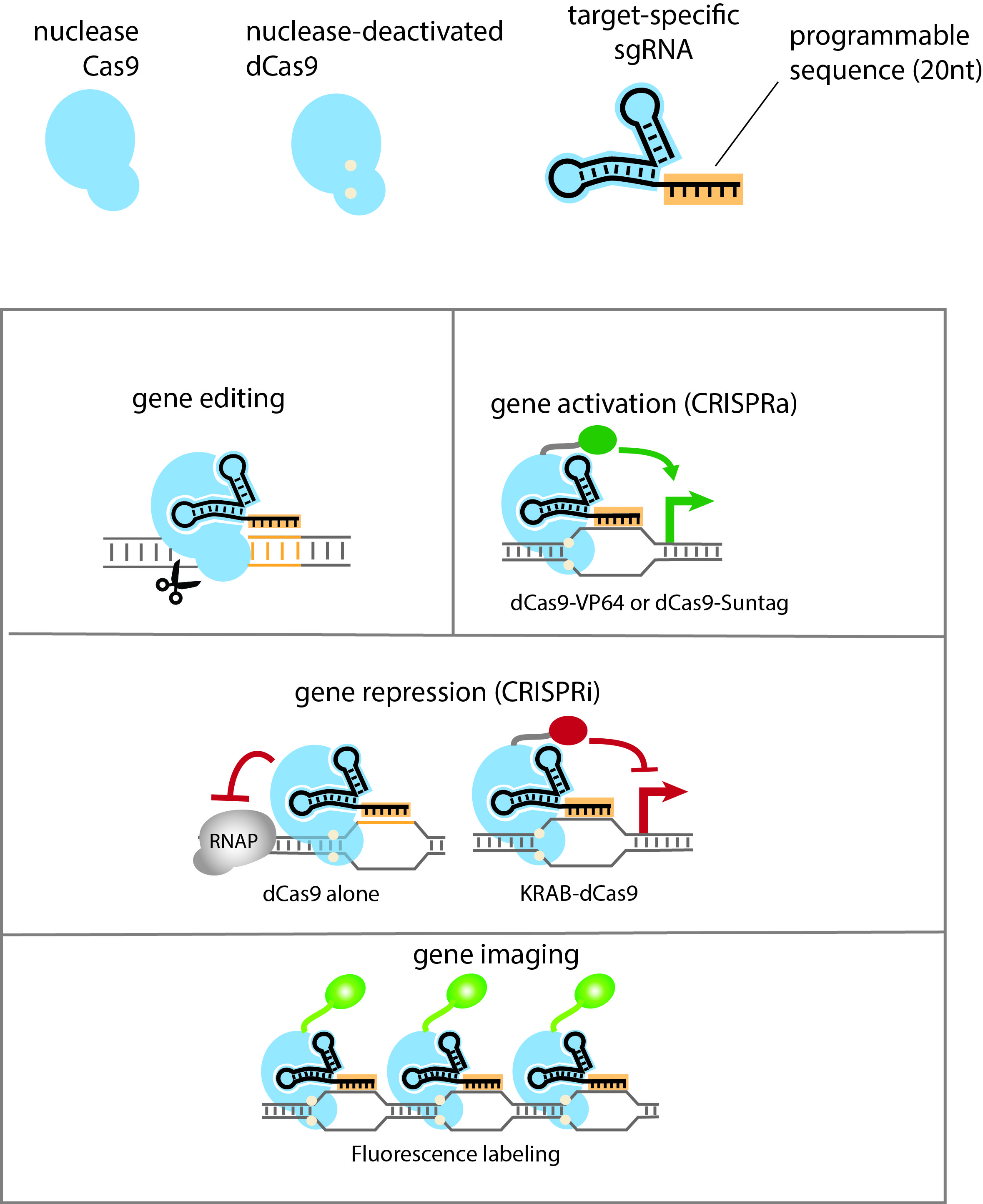
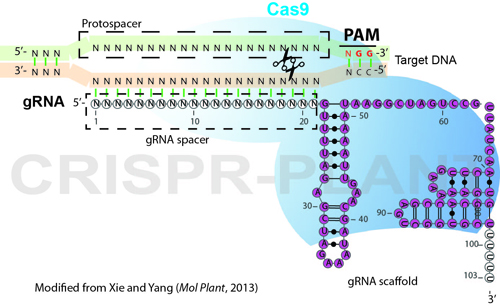
Identification of Spacer and Protospacer Sequence Requirements in the Vibrio cholerae Type I-E CRISPR/Cas System | mSphere
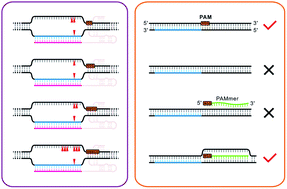
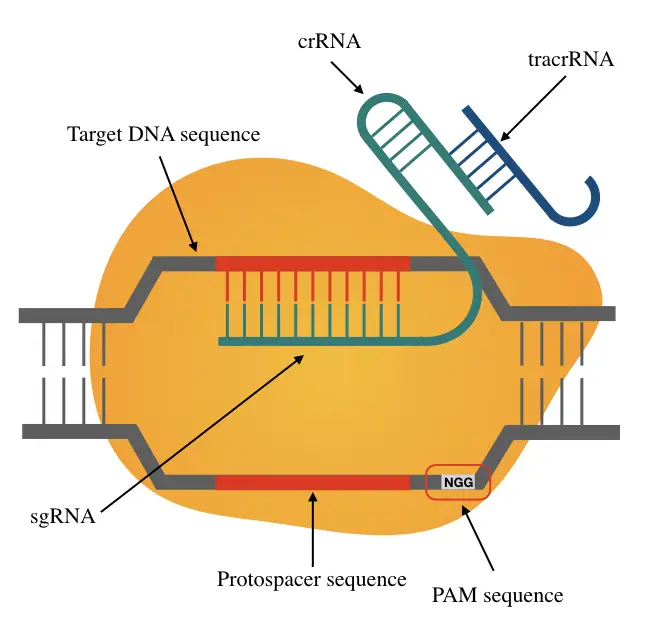
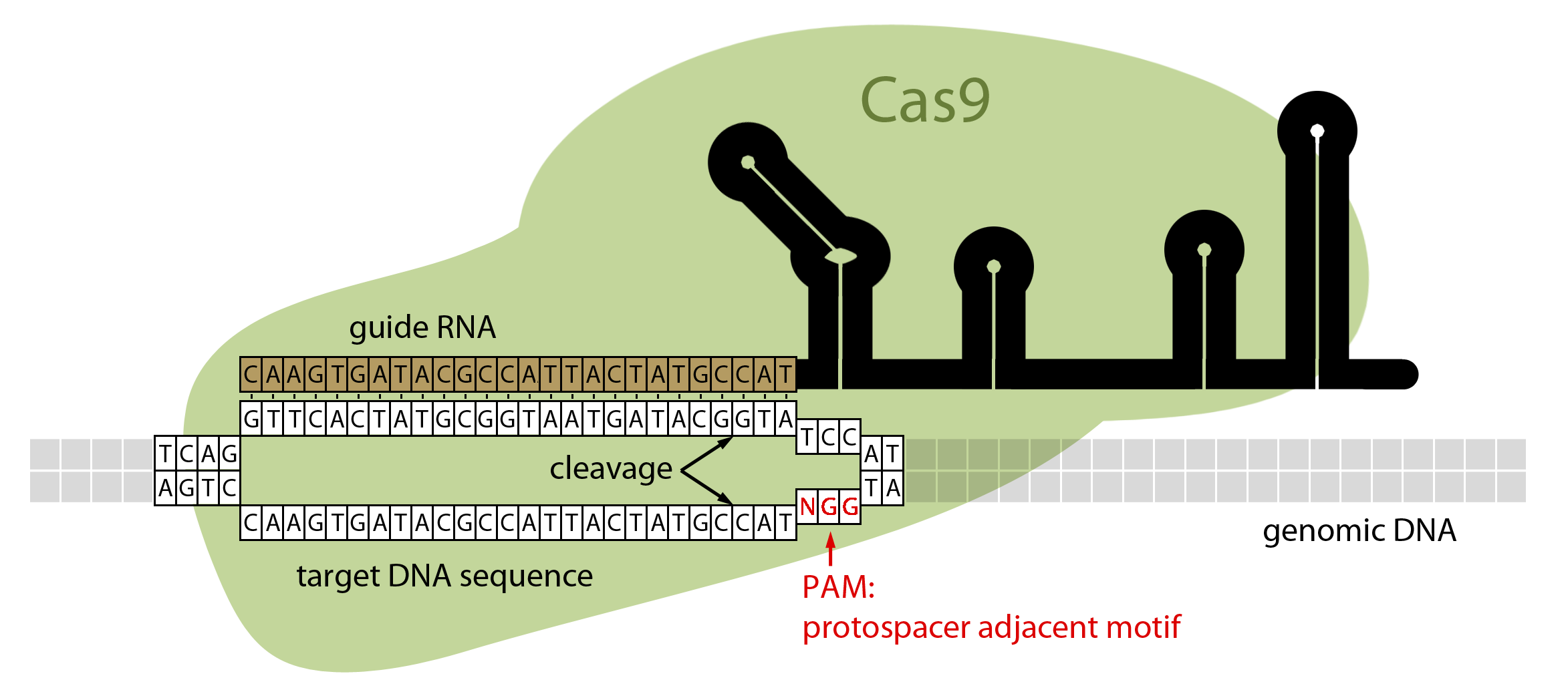
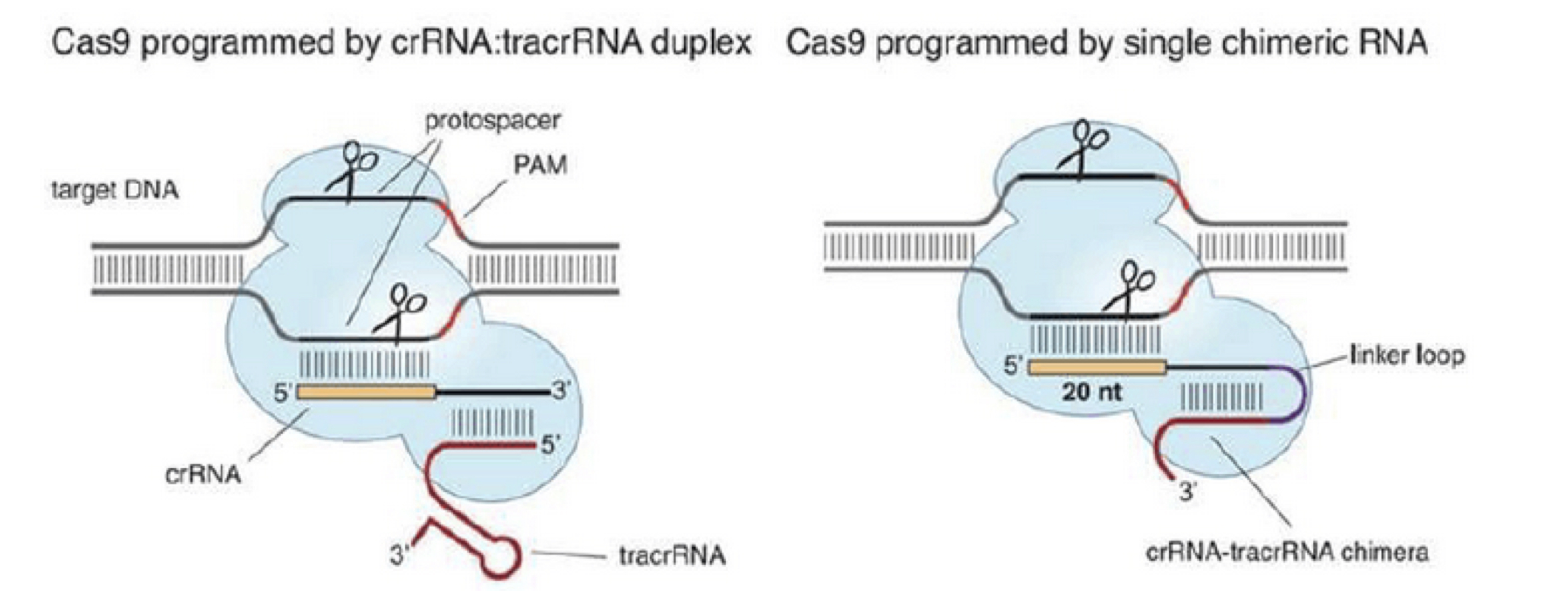
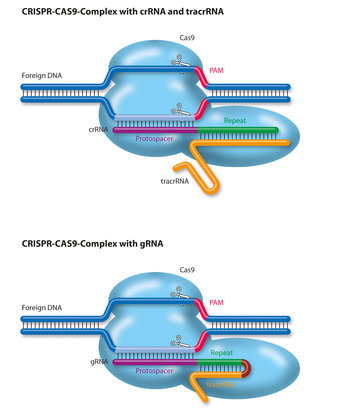
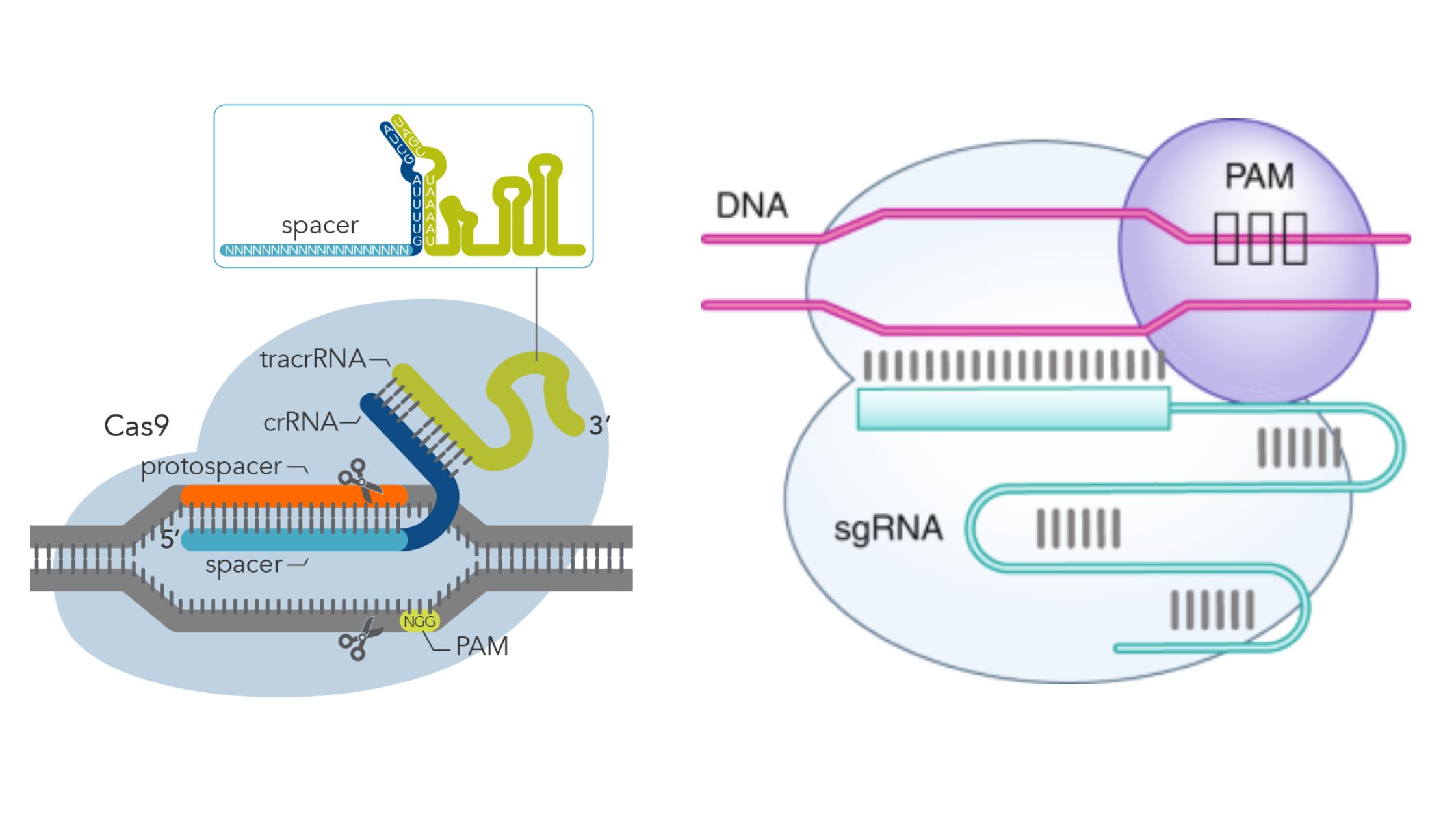
Scheme showing the functioning of CRISPR-Cas9 nuclease. PAM=protospacer... | Download Scientific Diagram
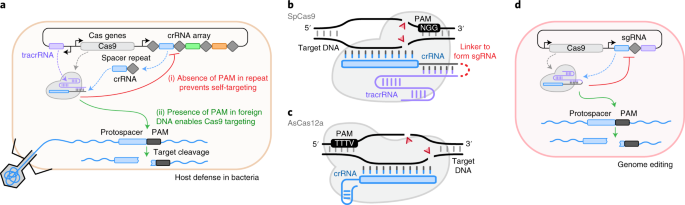
Scalable characterization of the PAM requirements of CRISPR–Cas enzymes using HT-PAMDA | Nature Protocols
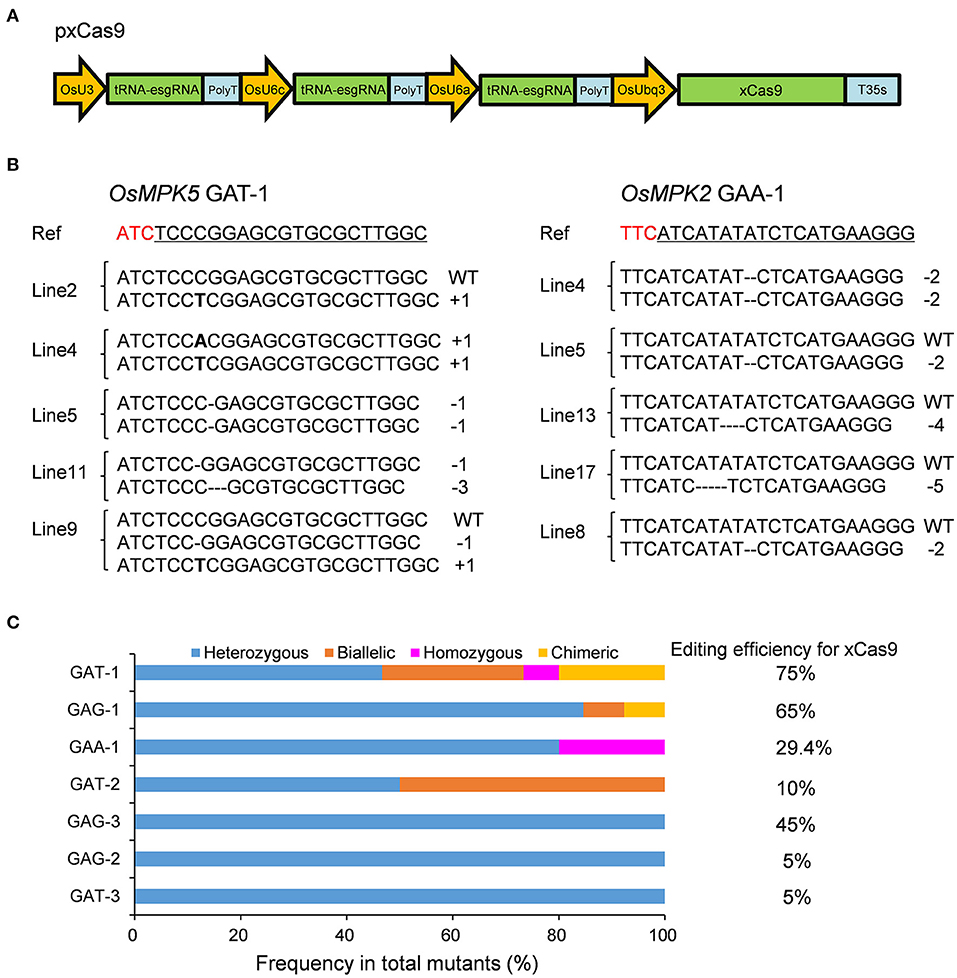
Frontiers | Genome Engineering in Plant Using an Efficient CRISPR-xCas9 Toolset With an Expanded PAM Compatibility
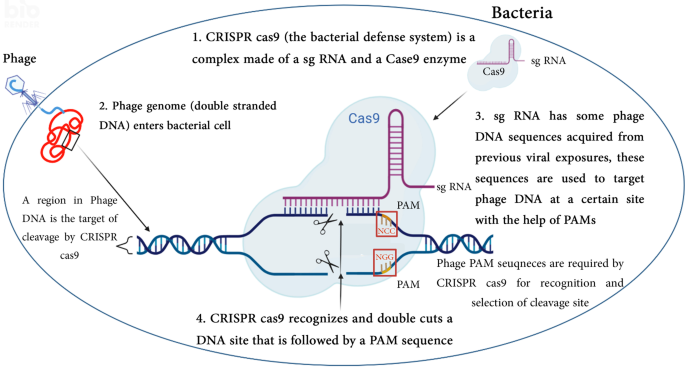
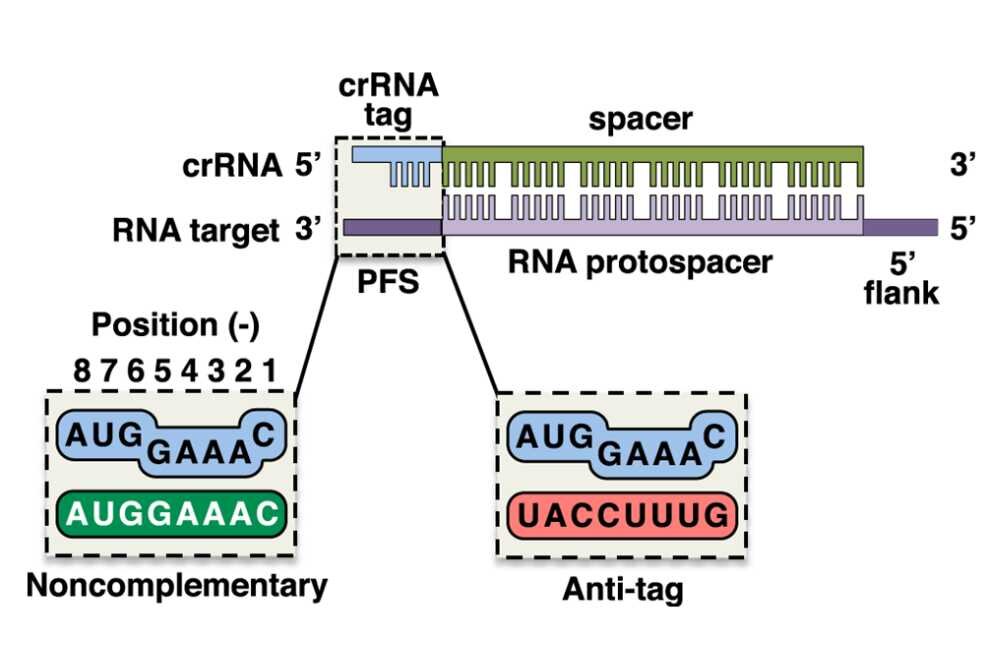
Bio-informatic analysis of CRISPR protospacer adjacent motifs (PAMs) in T4 genome | BMC Genomic Data | Full Text