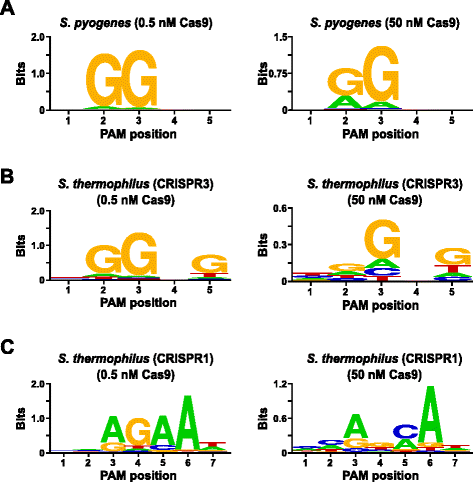
Rapid characterization of CRISPR-Cas9 protospacer adjacent motif sequence elements | Genome Biology | Full Text

In Silico Processing of the Complete CRISPR‐Cas Spacer Space for Identification of PAM Sequences - Mendoza - 2018 - Biotechnology Journal - Wiley Online Library

DNA topology regulates PAM-Cas9 interaction and DNA unwinding to enable near-PAMless cleavage by thermophilic Cas9 - ScienceDirect

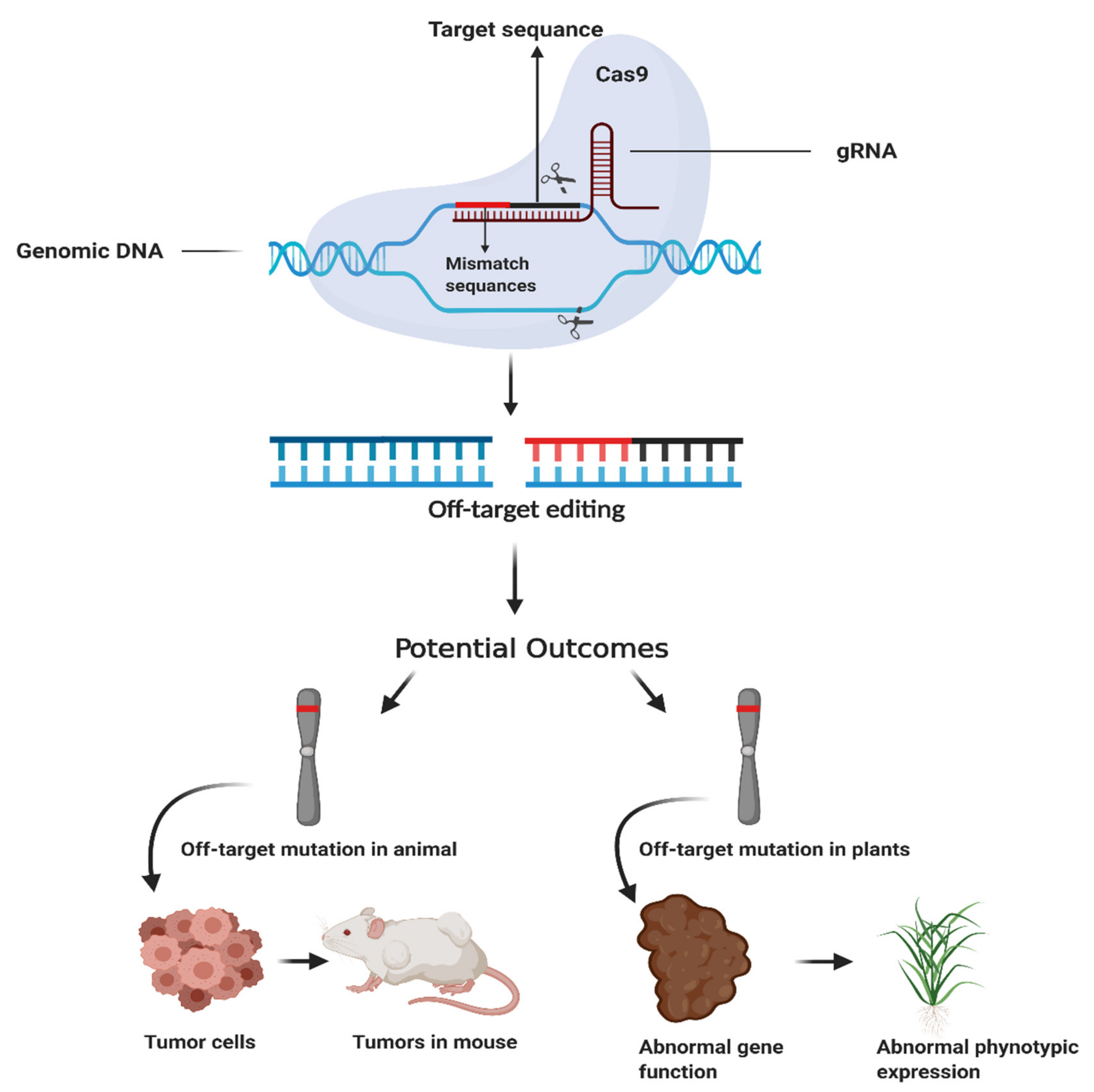
Nuclease Target Site Selection for Maximizing On-target Activity and Minimizing Off-target Effects in Genome Editing: Molecular Therapy

Schematic for identification of PAM preferences by Cas9 cleavage in... | Download Scientific Diagram

Comprehensive PAM prediction for CRISPR-Cas systems reveals evidence for spacer sharing, preferred strand targeting and conserved links with CRISPR repeats | bioRxiv

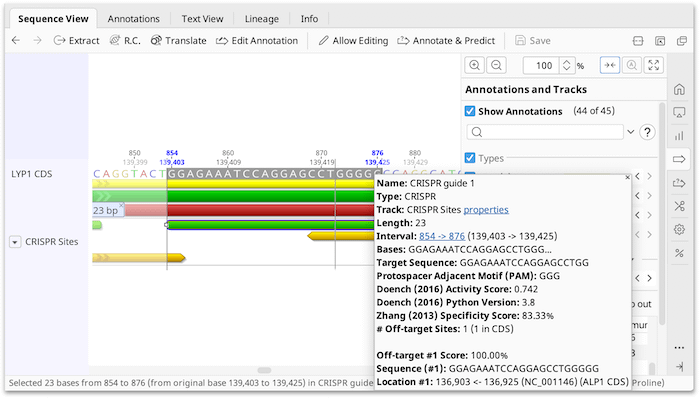
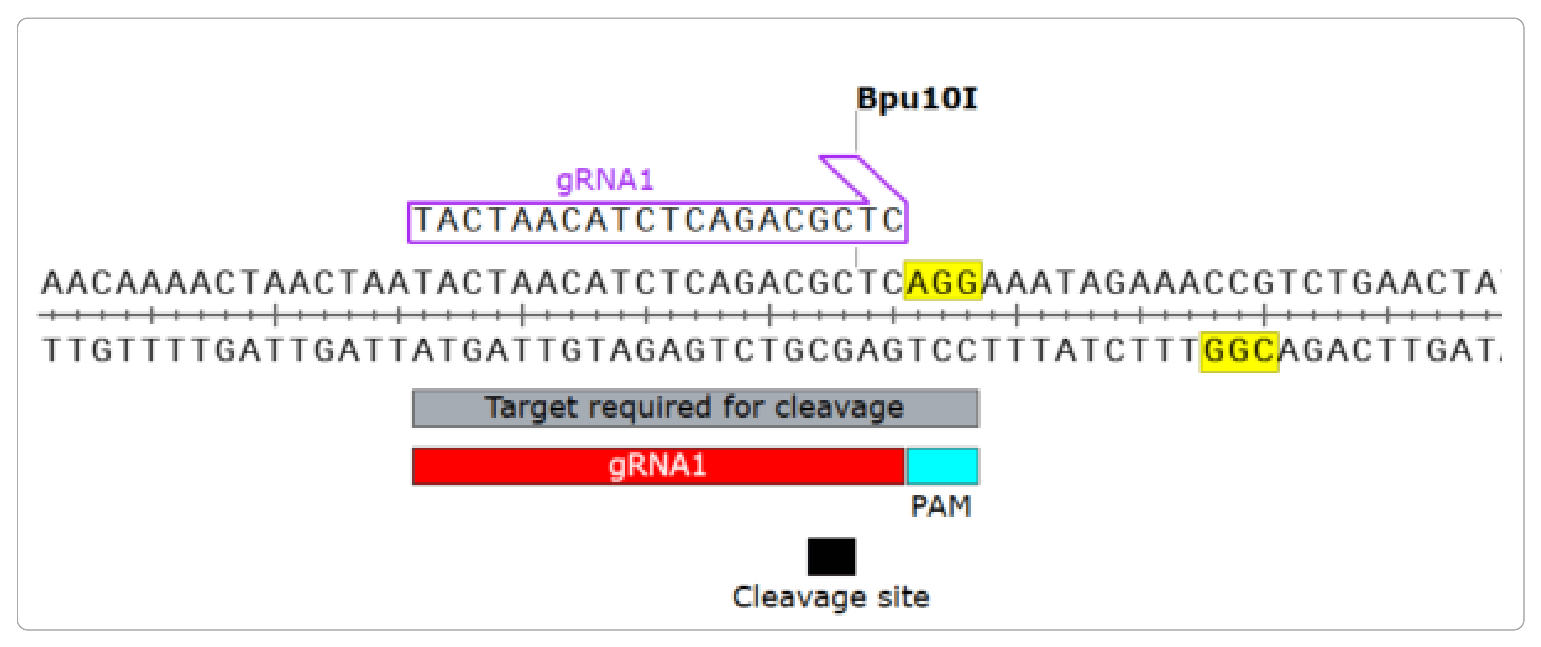
CT-Finder: A Web Service for CRISPR Optimal Target Prediction and Visualization | Scientific Reports

Identification of functional PAM sequences for CRISPR-interference.... | Download Scientific Diagram


.png?sfvrsn=58c63b07_0)