Convert your Fasta files in seconds! FASTA to multi-FASTA format converter. Merge FASTA files into a single multiFASTA. FASTA converter/merger
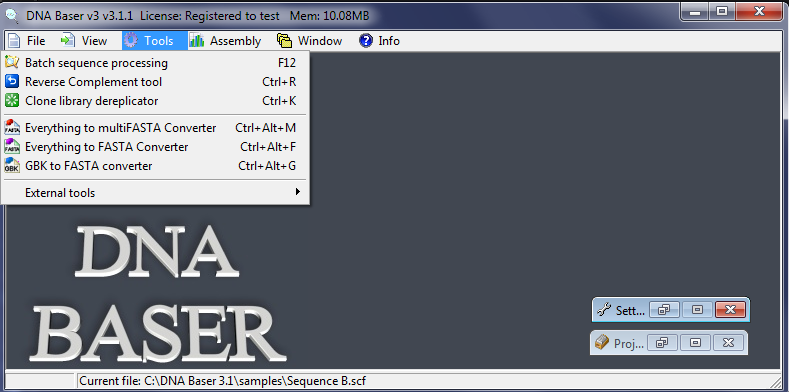
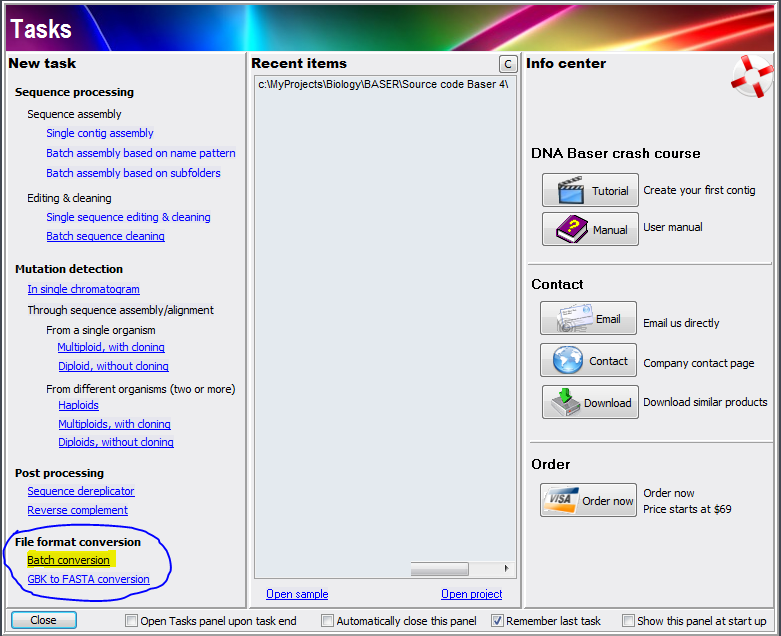
Everything to Fasta Converter converts the specified samples (SCF, ABI, FASTA, multiFasta, GBK, multiGBK, SEQ, TXT) to FASTA format. Converting any files to Fasta

Convert your Fasta files in seconds! FASTA to multi-FASTA format converter. Merge FASTA files into a single multiFASTA. FASTA converter/merger

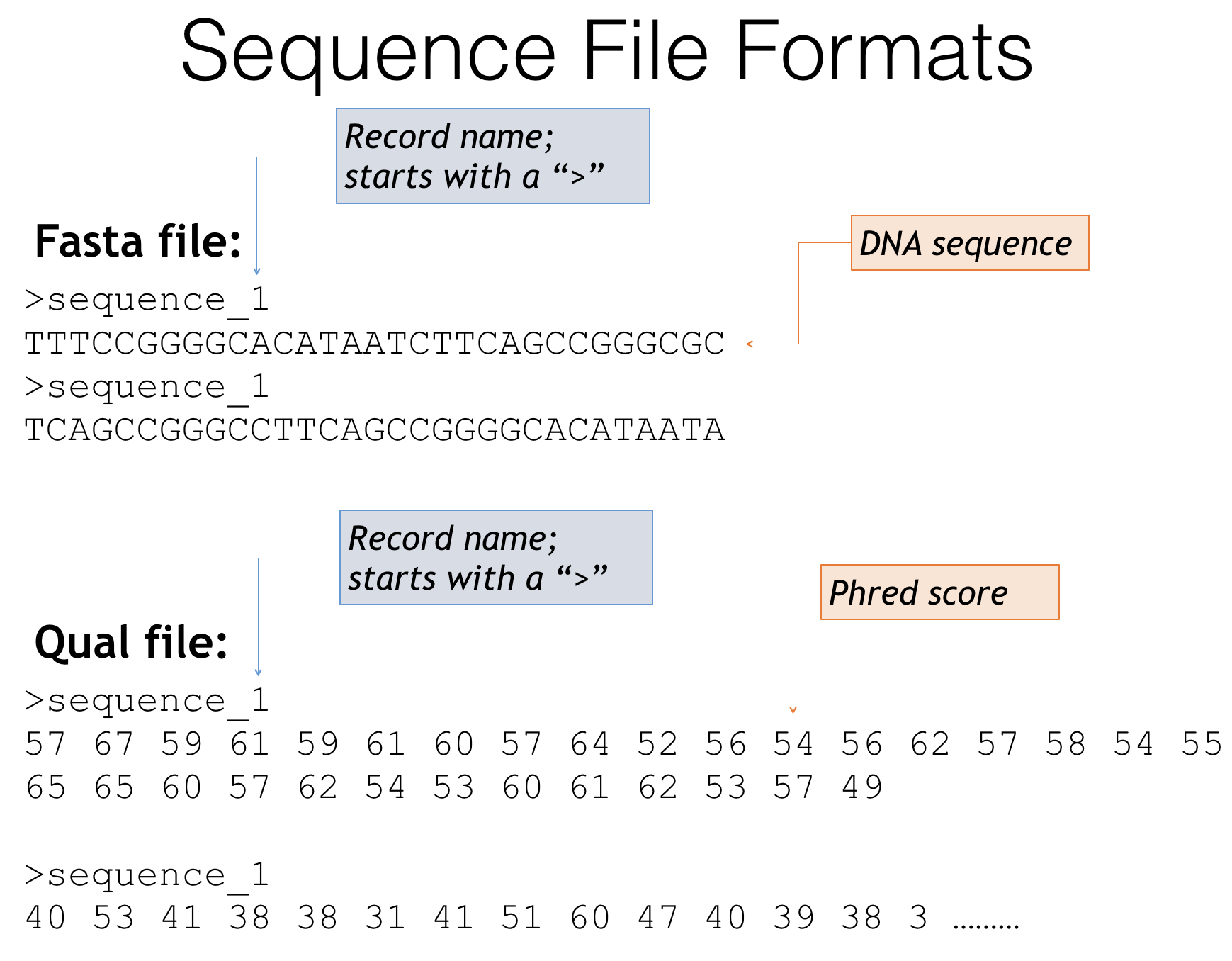
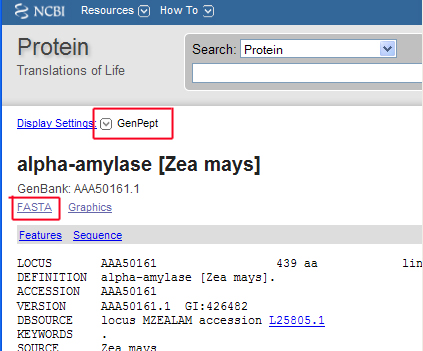
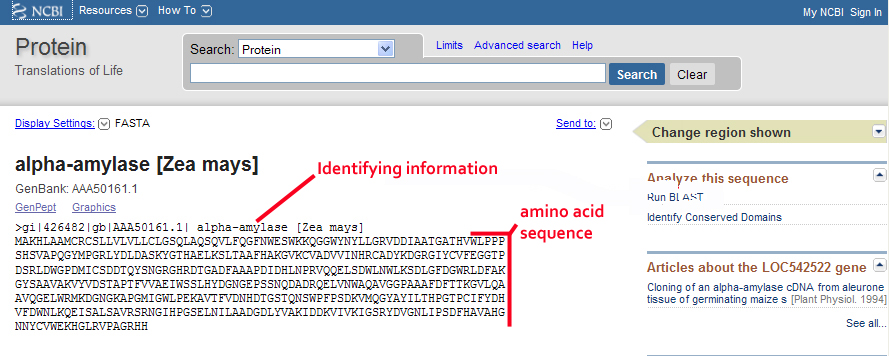
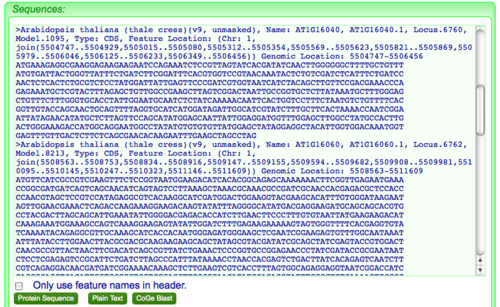
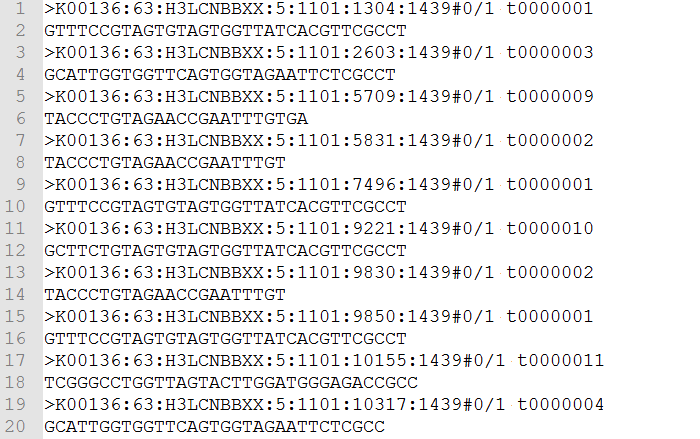
Aveyn's Blog: How to convert a sequence (c/DNA/RNA) into FASTA format quickly (the manual method)? and other handy tips in analyzing sequencing results.
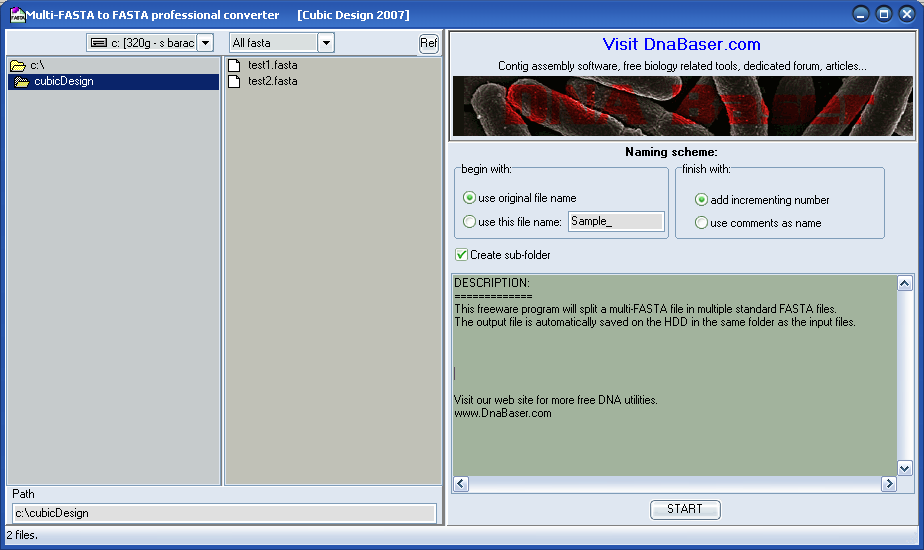
Everything to Fasta Converter converts the specified samples (SCF, ABI, FASTA, multiFasta, GBK, multiGBK, SEQ, TXT) to FASTA format. Converting any files to Fasta